Anatomical Priors for Image Segmentation via Post-Processing with Denoising Autoencoders
Introduction
The authors propose to use Denoising Autoencoders (DAE) as a post-processing step to impose shape constraints on the resulting masks obtained with arbitrary segmentation methods. They claim that Post-DAE can improve the quality of noisy and incorrect segmentation masks obtained with a variety of standard methods (CNN, RF-based classifiers, etc.) by bringing them back to a feasible space, with almost no extra computational time.
Prior works cited that tried to implement anatomical prior constraints include ACNN and connected conditional random field (CRF). These methods of applying priors are critized either for being used during training, and therefore not usable with any other segmentation method (e.g. ACNN), or for making the assumption that objects are usually continuous (e.g. CRF).
Methods
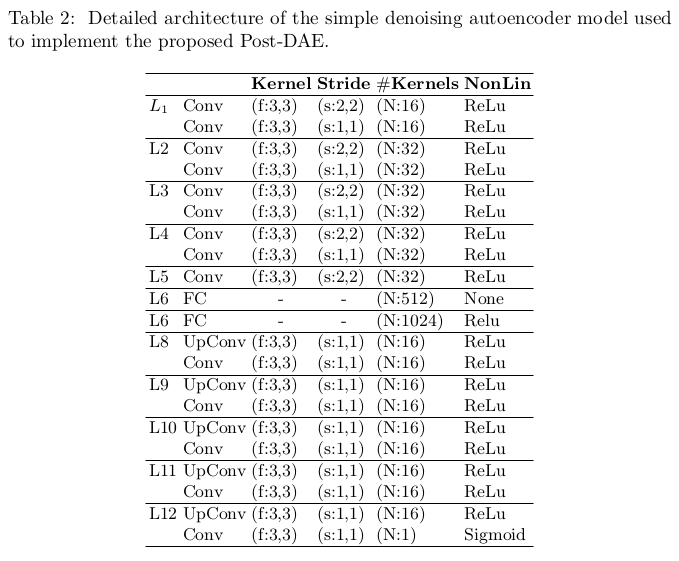
The autoencoder model used for denoising is a standard convolutional autoencoder with a latent space in 512 dimensions. The authors provided a full description of their model’s architecture (c.f. Fig.2). They also mention using dropout (\(p=0.5\)) for regularization after layer 5.
The ground truth masks were artificially degraded to produce the noisy masks used to train the DAE. The degradations were applied randomly from the following functions:
- addition and removal of random geometric shapes (circles, ellipses, lines and rectangles) to simulate over and under segmentations;
- morphological operations (e.g. erosion, dilation, etc) with variable kernels to perform more subtle mask modifications;
- random swapping of foreground-background labels in the pixels close to the mask borders.
The proposed method was not compared against method enforcing anatomical prior at train time, since the authors’ goal was to compare post-processing strategies. The post-processing method was benchmarked on two different segmentation methods that produce segmentation masks of various qualities.
Data
The method was benchmarked on lung segmentation in X-Ray images, using the Japanese Society of Radiological Technology (JSRT) database, which contains 247 PA chest X-ray images of 2048x2048 pixels and isotropic spacing of 0.175 mm/pixel. To train their DAE, the image were downsampled to 1024x1024 and divided 70/20/10 between training, validation and test sets. According to the authors, the lungs present “high variability among subjects”.
Results
Results show consistent improvement, especially in low quality segmentation masks. On this dataset, even well-trained models showcase holes in the lung and small isolated blobs. The post-processing significantly improves the Hausdorff distance in these cases, even for the well-trained models. The baseline post-processing method used for the comparison is a “classic fully-connected CRF”, but no further detail of its implementation is given.
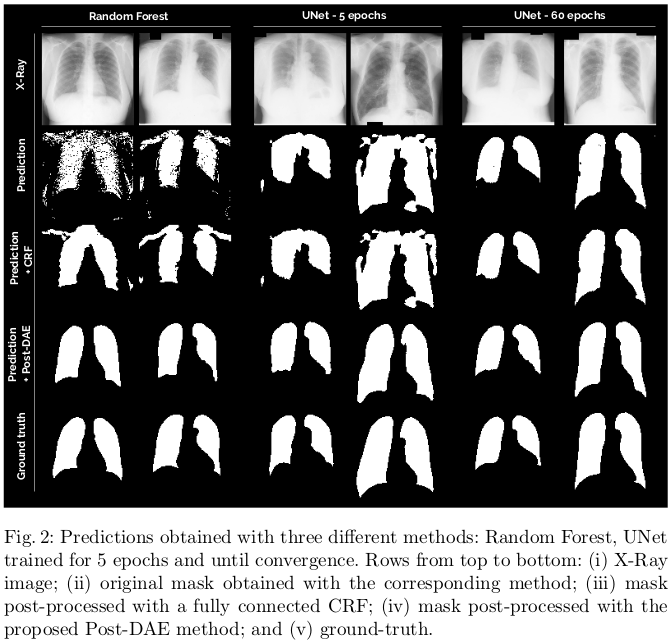
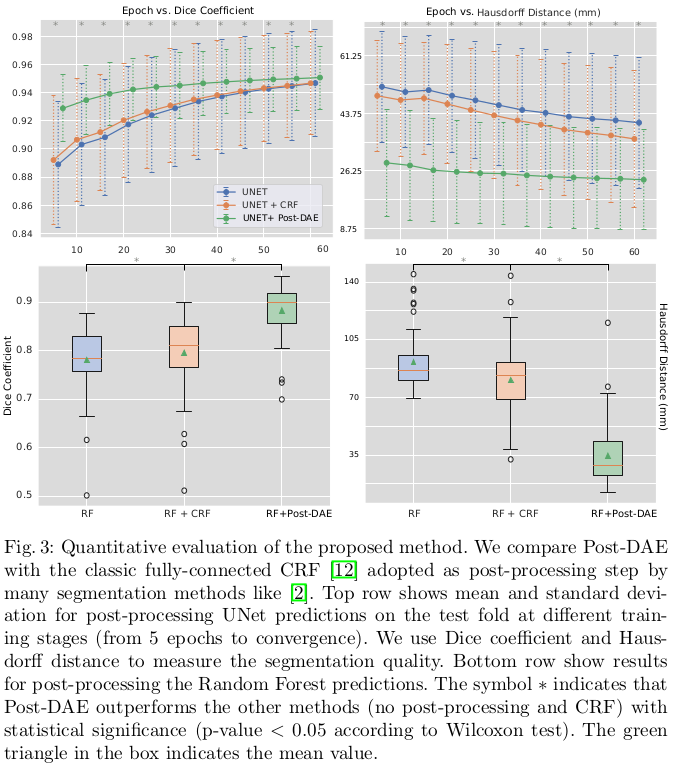
Conclusions
The authors admit themselves that “one of the limitations of Post-DAE is related to data regularity”. High variability still amounts to somehow uniform segmentations masks in terms of shape and topology, even in pathological cases. The authors mention that “cases like brain lesions or tumors where shape is not that regular” are out of the scope of the paper, but “will be explored as future work”, along with “multiclass and volumetric segmentation cases”.