Towards increased trustworthiness of deep learning segmentation methods on cardiac MRI
Summary
In this paper, the authors explore ways for estimating the trustworthiness of segmentation results obtained with a CNN. They tested their methods on the ACDC MRI Cardiac dataset. One motivation for this paper is the fact that state-of-the-art methods often “locally produce anatomically implausible results that medical experts would not make”. Since the proposed method implies “human-in-the-loop”, it is semi-automatic.
Proposed method
They implemented a Dilated CNN with three losses : soft-Dice (SD), cross-entropy (CE), and the Brier score (BS) (which is equal to the average gap between softmax probabilities and the references).
They then tested two types of uncertainty maps
- entropy maps (e-maps): which is the per-pixel multi-class entropy
- Bayesian uncertainty maps (u-maps): which is obtained with a Monte-Carlo dropout.
Results
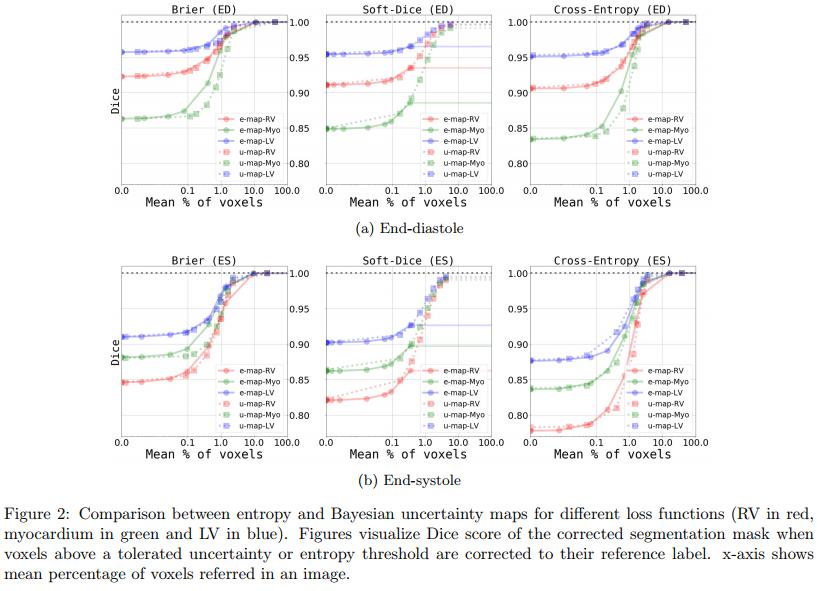
Results in Figure 2 show the two main conclusions of the paper :
- The Brier loss is better than the other 2 losses
- e-maps and u-maps are similar when replacing voxels whose uncertainty is above a certain threshold by their GT value.
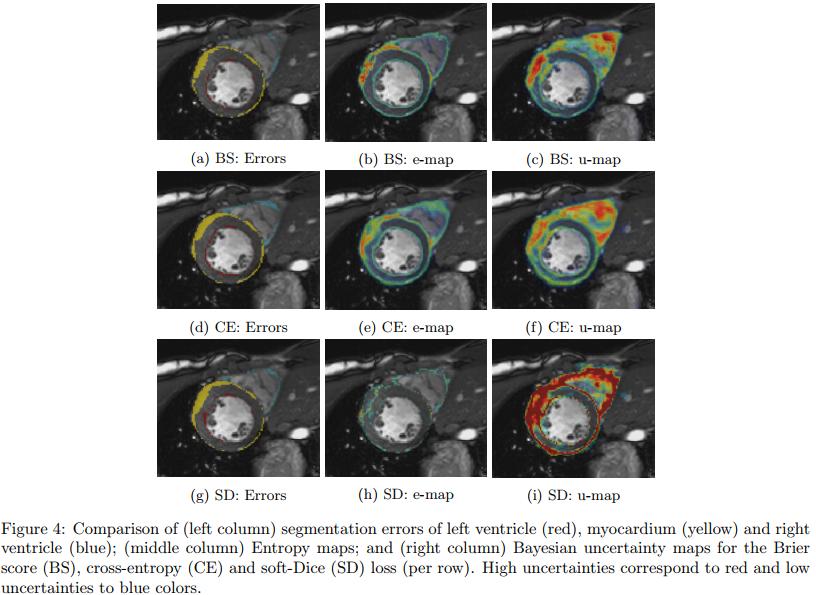
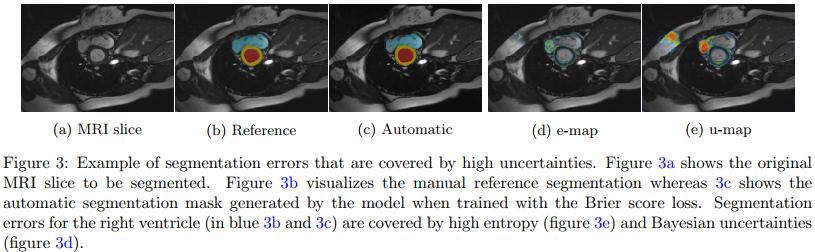
Figure 3 and 4 show that the uncertainty maps are close to the reported segmentation errors although not perfect. Unfortunately, the authors do not really mention false positive uncertainties and do not perform any experiments with real end-users.